白皮杉醇調控miR-106b-5p/RUNX3軸對宮頸癌細胞遷移及侵襲的影響
王玉寧 宋聚星 田志剛 郝國榮 申浩


摘要:目的 探究白皮杉醇(PIC)調控微小RNA-106b-5p(miR-106b-5p)/RUNT相關轉錄因子3(RUNX3)軸對宮頸癌(CC)細胞遷移及侵襲的影響。方法 使用不同濃度PIC(0、20、40、80和160 μmol/L)培養液處理人CC Hela細胞,通過CCK-8法檢測PIC對細胞增殖活力的影響,以確定PIC最佳使用濃度。將Hela細胞分為Control組、PIC組、PIC+NC mimics組、PIC+miR-106b-5p mimics組、NC inhibitor組、miR-106b-5p inhibitor組、miR-106b-5p inhibitor+si-RNA組及miR-106b-5p inhibitor+si-RUNX3組。實時熒光定量PCR檢測各組Hela細胞miR-106b-5p表達水平;Transwell法檢測各組Hela細胞遷移及侵襲能力;Western blot法檢測各組Hela細胞中RUNX3、基質金屬蛋白酶(MMP)2和MMP9蛋白表達水平;雙螢光素酶報告基因實驗檢測miR-106b-5p與RUNX3靶向關系。結果 Hela細胞增殖活力隨著PIC處理濃度的增高而呈現降低趨勢(P<0.05),其中80 μmol/L PIC對Hela細胞的抑制作用接近半數抑制濃度(IC50),故選擇80 μmol/L為后續研究的PIC濃度。與Control組相比,PIC組miR-106b-5p、MMP2和MMP9表達水平均降低,遷移和侵襲細胞數量減少,RUNX3表達水平增加(P<0.05);與PIC+NC mimics組相比,PIC+miR-106b-5p mimics組miR-106b-5p、MMP2和MMP9表達水平增加,遷移和侵襲細胞數量增多,RUNX3表達水平降低(P<0.05)。雙螢光素酶報告基因實驗證實RUNX3為miR-106b-5p的靶基因。與NC inhibitor組相比,miR-106b-5p inhibitor組RUNX3表達水平增加,miR-106b-5p、MMP2與MMP9表達水平降低,遷移和侵襲細胞數量減少(P<0.05);與miR-106b-5p inhibitor+si-RNA組相比,miR-106b-5p inhibitor+si-RUNX3組RUNX3表達水平降低,miR-106b-5p、MMP2和MMP9表達水平增加,遷移和侵襲細胞數量增多(P<0.05)。結論 PIC通過抑制miR-106b-5p表達、促進RUNX3表達來抑制CC細胞遷移及侵襲。
關鍵詞:微RNAs;核心結合因子α3亞基;宮頸腫瘤;細胞運動;腫瘤浸潤;白皮杉醇;miR-106b-5p;RUNT相關轉錄因子3
中圖分類號:R285,R349.5文獻標志碼:ADOI:10.11958/20221529
The effect of piceatannol on the migration and invasion of cervical cancer cells by regulating miR-106b-5p/RUNX3 axis
WANG Yuning SONG Juxing TIAN Zhigang HAO Guorong SHEN Hao
1 Department of Gynecology, Shijiazhuang No.4 Hospital, Shijiazhuang 050035, China; 2 Department of Pathology,
3 Department of Clinical Laboratory, Baoding Second Central Hospital
Abstract: Objective To explore the effect of piceatannol (PIC) on migration and invasion of cervical cancer (CC) cells by regulating microRNA-106b-5p (miR-106b-5p)/RUNT-related transcription factor 3 (RUNX3) axis. Methods Firstly, human CC cells Hela were treated with culture media of different concentrations of PIC (0, 20, 40, 80 and 160 μmol/L), and the effect of PIC on cell proliferation was detected by CCK-8 assay to determine the optimal concentration of PIC. Hela cells were divided into the control group, the PIC group, the PIC+NC mimics group, the PIC+ miR-106b-5p mimics group, the NC inhibitor group, the miR-106b-5p inhibitor group, the miR-106b-5p inhibitor+si-RNA group and the miR-106b-5p inhibitor+si-RUNX3 group. qRT-PCR was used to detect the expression level of miR-106b-5p in Hela cells in each group. Transwell method was used to detect the migration and invasion abilities of Hela cells in each group. Western blot assay was used to detect the protein levels of RUNX3, MMP2 and MMP9 in Hela cells of each group. Dual luciferase reporter gene experiment was used to detect the targeting relationship between miR-106b-5p and RUNX3. Results The proliferation activity of Hela cells decreased with the increase of PIC treatment concentration (P<0.05). The inhibitory effect of 80 μmol/L PIC on Hela cells was nearly to half inhibitory concentration (IC50), so 80 μmol/L was selected as the PIC concentration for subsequent study. Compared with the control group, the expression levels of miR-106b-5p, MMP2 and MMP9 decreased in the PIC group, the numbers of migrating and invasive cells decreased, and the expression level of RUNX3 increased (P<0.05). Compared with the PIC+NC mimics group, the expression levels of miR-106b-5p, MMP2 and MMP9 increased in the PIC+miR-106b-5p mimics group, numbers of migrating and invasive cells increased, and the expression level of RUNX3 decreased (P<0.05). Double luciferase reporter gene assay confirmed RUNX3 as the target gene of miR-106b-5p. Compared with the NC inhibitor group, the expression level of RUNX3? increased in the miR-106b-5p inhibitor group, and the expression levels of miR-106b-5p, MMP2 and MMP9 decreased, and numbers of migrating and invasive cells decreased (P<0.05). Compared with the miR-106b-5p inhibitor+si-RNA group, the miR-106b-5p inhibitor+si-RUNX3 group showed lower RUNX3 expression level, increased miR-106b-5p, MMP2 and MMP9 expression levels, and increased numbers of migrating and invading cells (P<0.05). Conclusion PIC inhibits the miR-106b-5p expression and promotes RUNX3 expression to inhibit CC cell migration and invasion.
Key words: microRNAs; core binding factor alpha 3 subunit; uterine cervical neoplasms; cell movement; neoplasm invasiveness; piceatannol; miRNA-106b-5p; RUNT-related transcription factor 3
宮頸癌(cervical cancer,CC)是一種常見的女性惡性腫瘤,并且是全球癌癥相關死亡的主要原因之一[1]。由于其病因復雜,對化療藥物的耐藥性和高轉移潛力,晚期CC患者預后較差[2-3]。因此,迫切需要開發一種高效、低毒的新型CC治療藥物。白皮杉醇(piceatannol,PIC)是一種酚類化合物和白藜蘆醇的類似物,具有抗炎、抗氧化、調節免疫等多種藥理特性[4],且對不同類型的腫瘤,如骨肉瘤[5]、膀胱癌[6]、結腸癌[7]具有抗癌活性。然而,其在CC中的作用和分子機制尚不清楚。微小RNAs(miRNAs)的發現為癌細胞轉移和進展的分子機制研究提供了新的方法和見解。有研究發現,miR-106b在CC組織中高表達,可能與CC的發展有關[8]。miRNAs可通過調控靶基因的翻譯或降解參與細胞增殖、凋亡、遷移及侵襲等多種過程。如在肝癌中,miR-106b-5p通過靶向下調RUNT相關轉錄因子3(RUNT-related transcription factor 3,RUNX3)表達促進腫瘤細胞的侵襲[9]。已知RUNX3在CC組織中低表達,為抑癌基因[10]。本研究旨在探究PIC對CC細胞增殖、遷移及侵襲的影響,并分析其潛在機制,以期為尋找新型CC化療藥物提供參考。
1 材料與方法
1.1 實驗材料
人CC Hela細胞購于中國科學院細胞庫;DMEM培養基、胎牛血清、青/鏈霉素、LipofectamineTM 2000轉染試劑均購自美國Invitrogen公司。PIC購自成都麥得生物科技公司,純度>99%。miR-106b-5p模擬物(miR-106b-5p mimics)和其陰性對照(NC mimics)、miR-106b-5p抑制物(miR-106b-5p inhibitor)和其陰性對照(NC inhibitor)、RUNX3小干擾RNA(si-RUNX3)和其陰性對照(si-RNA)、野生型RUNX3(RUNX3-WT)和突變型RUNX3(RUNX3-MUT)及miR-106b-5p和U6引物均由生工生物工程(上海)股份有限公司設計合成。CCK-8細胞增殖檢測試劑盒購自日本Dojindo;細胞總RNA提取試劑盒(Trizol法)、逆轉錄試劑盒、實時熒光定量聚合酶鏈反應(qPCR)試劑盒購自南京Vazyme公司。Transwell小室、基質膠購自美國Corning公司;高效RIPA細胞快速裂解液購自北京Solarbio公司;兔源RUNX3、基質金屬蛋白酶(MMP)2、MMP9、甘油醛-3-磷酸脫氫酶(GAPDH)抗體均購自英國Abcam公司;雙螢光素酶活性檢測試劑盒購自北京百奧萊博科技有限公司。HERAcell 240i型二氧化碳細胞培養箱、Varioskan LUX型酶標儀購自美國Thermo Fisher公司;CFX96型熒光定量PCR儀、OI100 Touch型凝膠成像系統購自美國Bio-Rad公司;CKX53型倒置顯微鏡購自日本OLYMPUS公司。
1.2 研究方法
1.2.1 細胞培養與分組
Hela細胞接種在含有10%胎牛血清和1%青/鏈霉素的DMEM培養基,置于37 ℃、5%CO2培養箱中培養。取對數生長期的Hela細胞進行實驗,首先采用含不同濃度PIC(0、20、40、80和160 μmol/L)的培養液處理Hela細胞24 h,通過CCK-8法檢測不同濃度的PIC對Hela細胞增殖活力的影響,以確定最佳PIC使用濃度。將細胞分為:Control組(正常培養Hela細胞)、PIC組(含有80 μmol/L PIC的培養基處理)、PIC+NC mimics組(轉染NC mimics后采用80 μmol/L PIC處理)、PIC+miR-106b-5p mimics組(轉染miR-106b-5p mimics后采用80 μmol/L PIC處理)、NC inhibitor組(轉染NC inhibitor)、miR-106b-5p inhibitor組(轉染miR-106b-5p inhibitor)、miR-106b-5p inhibitor+si-RNA組(轉染miR-106b-5p inhibitor和si-RNA)及miR-106b-5p inhibitor+si-RUNX3組(轉染miR-106b-5p inhibitor和si-RUNX3),細胞轉染嚴格遵循LipofectamineTM 2000說明書流程。
1.2.2 CCK-8檢測不同濃度PIC處理后的Hela細胞增殖活力
將對數生長期的Hela細胞以1×104個/孔的密度接種于96孔板中,采用含不同濃度PIC(0、20、40、80和160 μmol/L)培養液處理Hela細胞24 h,每孔中添加10 μL CCK-8溶液混勻后繼續培養4 h,在酶標儀中測定450 nm波長處的吸光度(A)值,計算細胞增殖活力。細胞增殖活力(%)=APIC加藥組/APIC 0 μmol/L組×100%,確定PIC最佳使用濃度。
1.2.3 qPCR檢測各組Hela細胞中miR-106b-5p表達水平
參照1.2.1中分組方法處理Hela細胞后,棄去培養基,使用Trizol試劑提取各組細胞總RNA,將RNA通過逆轉錄試劑盒逆轉錄為cDNA,隨后采用qPCR試劑盒進行PCR擴增。miR-106b-5p引物:上游5′-TGCGGCAACACCAGTCGATGG-3′,下游5′-CCAGTGCAGGGTCCGAGGT-3′;內參U6引物:上游5′-CTCGCTTCGGGCAGCACA-3′,下游5′-AACGCTTCACGAATTTGCGT-3′。反應體系:2 ?L cDNA(200 μg/L),上、下游引物(10 ?mol/L)各0.8 ?L,10 ?L SYBR Premix Ex Taq(2×),6.4 ?L ddH2O。反應條件:95 ℃預變性30 s;95 ℃變性5 s,55 ℃退火10 s,72 ℃延伸15 s,共40個循環。以U6為內參基因,使用2-ΔΔct法計算miR-106b-5p的相對表達水平。
1.2.4 Transwell法檢測各組Hela細胞遷移及侵襲能力
參照1.2.1中的分組方法處理Hela細胞后,棄去培養基,使用無血清的培養基懸浮細胞,吸取200 μL細胞懸浮液(1×106個/mL)加入Transwell小室上室中,下室中添加500 μL正常培養基。培養24 h后,取出小室,拭去上室中未遷移的細胞。4%多聚甲醛固定10 min,1%結晶紫染色5 min,晾干后于倒置顯微鏡下拍照記錄。隨機對5個視野細胞進行計數,取均值表示遷移細胞數量。侵襲實驗將Transwell小室提前用基質膠包被后使用,其余步驟同上。
1.2.5 Western blot檢測各組Hela細胞中RUNX3、MMP2和MMP9蛋白水平
參照1.2.1中分組方法處理Hela細胞后,棄去培養基,使用RIPA法提取各組細胞的總蛋白,并通過Bradford法定量。通過十二烷基硫酸鈉聚丙烯酰胺凝膠電泳分離蛋白質并轉移到PVDF膜上。脫脂牛奶封閉1 h,添加一抗RUNX3(1︰500)、MMP2(1︰1 000)、MMP9(1︰1 000)及內參GAPDH(1︰2 000),在4 ℃下孵育過夜。隔日使用辣根過氧化物酶偶聯的二抗(1︰4 000)在室溫下孵育2 h。最后使用凝膠成像系統拍照記錄蛋白印跡,使用Image J分析蛋白條帶的灰度值,量化蛋白表達數據。目的蛋白的相對表達量=目的蛋白的灰度值/內參蛋白的灰度值。
1.2.6 雙螢光素酶報告基因實驗檢測miR-106b-5p與RUNX3靶向關系
將Hela細胞以1×105個/孔的密度接種于24孔細胞培養板中,分別使用RUNX3-WT或RUNX3-MUT和miR-106b-5p mimics或NC mimics共同轉染24 h。使用雙螢光素酶活性檢測試劑盒檢測各組Hela細胞熒光素酶活性。
1.3 統計學方法
采用SPSS 25.0進行數據分析。符合正態分布的計量資料以x±s表示,使用單因素方差分析比較多組間的差異,組間多重比較使用SNK-q檢驗。P<0.05為差異有統計學意義。
2 結果
2.1 不同濃度PIC對Hela細胞增殖活力的影響
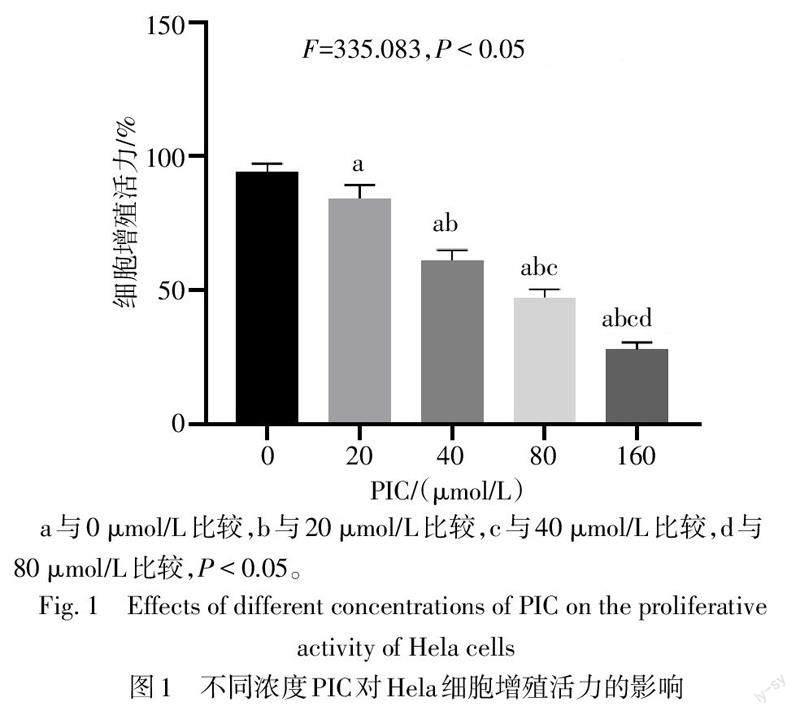
與0 μmol/L相比,20、40、80和160 μmol/L PIC處理后Hela細胞增殖活力降低(P<0.05),且隨著PIC濃度的增高,其對Hela細胞增殖活力的抑制作用也隨之增強,見圖1。PIC對Hela細胞增殖活力的半數抑制濃度(IC50)為75.858 μmol/L,因80 μmol/L PIC對Hela細胞的抑制率接近IC50,故選擇此濃度作為后續研究的PIC濃度。
2.2 過表達miR-106b-5p對PIC抑制Hela細胞遷移及侵襲和促進RUNX3表達的影響
與Control組相比,PIC組miR-106b-5p、MMP2和MMP9表達水平均降低,遷移和侵襲細胞數量減少,RUNX3表達水平增加(P<0.05);與PIC+NC mimics組相比,PIC+miR-106b-5p mimics組miR-106b-5p、MMP2和MMP9表達水平均增加,遷移和侵襲細胞數量增多,RUNX3表達水平降低(P<0.05);而PIC+NC mimics組與PIC組細胞中miR-106b-5p、RUNX3、MMP2、MMP9表達水平及遷移和侵襲細胞數量差異均無統計學意義(P>0.05),見表1,圖2、3。
2.3 miR-106b-5p與RUNX3靶向關系驗證
經miRTarBase網站預測(http://mirtarbase.mbc.nctu. edu.tw/php/index.php)發現,miR-106b-5p與RUNX3存在靶向結合位點,見圖4。雙螢光素酶實驗結果顯示,上調miR-106b-5p表達可有效降低轉染RUNX3-WT后的細胞相對螢光素酶活性(P<0.05),而對轉染RUNX3-MUT后的細胞相對螢光素酶活性則無明顯影響(P>0.05),見表2。
2.4 RUNX3對miR-106b-5p調控Hela細胞遷移及侵襲的影響
與NC inhibitor組相比,miR-106b-5p inhibitor組RUNX3表達水平增加,miR-106b-5p、MMP2與MMP9表達水平均降低,遷移和侵襲細胞數量減少(P<0.05);與miR-106b-5p inhibitor+si-RNA組相比,miR-106b-5p inhibitor+si-RUNX3組RUNX3表達水平降低,miR-106b-5p、MMP2和MMP9表達水平均增加,遷移和侵襲細胞數量增多(P<0.05);而miR-106b-5p inhibitor+si-RNA組與miR-106b-5p inhibitor組細胞中miR-106b-5p、RUNX3、MMP2與MMP9表達水平及遷移和侵襲細胞數量差異均無統計學意義(P>0.05),見表3,圖5、6。
3 討論
白藜蘆醇通過抑制細胞生長在CC中發揮抗癌作用[11],而PIC是一種天然的白藜蘆醇類似物。Wang等[5]研究發現,PIC以劑量依賴性方式抑制骨肉瘤細胞增殖,并誘導骨肉瘤細胞凋亡。Li等[6]發現PIC以濃度和時間依賴性方式抑制膀胱癌細胞的增殖,誘導細胞凋亡和細胞周期停滯。本研究發現,PIC以劑量依賴性方式抑制Hela細胞增殖,且PIC可降低Hela細胞的遷移和侵襲能力。這些數據表明PIC很有可能是CC的潛在治療劑。
miRNAs可以通過靶向調控其靶基因來參與腫瘤細胞增殖、遷移及侵襲,發揮抑癌基因或促癌基因的作用[12]。以往研究表明,白藜蘆醇及其類似物可調控miRNA表達來介導前列腺癌的治療[13]。Du等[14]研究發現,PIC通過上調miR-181a的表達誘導黑色素瘤細胞凋亡。因此,筆者推測PIC可能通過介導miRNA對CC細胞發揮抗癌作用。本研究發現PIC可抑制miR-106b-5p表達。已知miR-106b-5p在結直腸癌[15]、肝癌[16]等腫瘤組織及細胞中高表達,可通過調控其靶基因發揮促癌作用。為了進一步研究PIC的抗癌作用是否受miR-106b-5p表達的調節,本研究在上調miR-106b-5p的同時使用PIC處理Hela細胞,發現上調miR-106b-5p可逆轉PIC的抗腫瘤效應,提示PIC可能通過下調miR-106b-5p表達對CC細胞發揮抗癌作用。然而,其潛在分子機制還需進一步研究。
本研究通過在線網站預測發現,RUNX3存在miR-106b-5p靶向結合位點。Gu等[9]報道,miR-106b-5p通過靶向調控RUNX3表達促進肝癌細胞的發展。研究顯示,RUNX3作為TGF-β介導信號通路的重要轉錄因子發揮作用,在多種癌癥中發揮抑癌作用[17]。Song等[18]發現,RUNX3在胃癌組織和細胞中下調,上調其表達可抑制癌細胞的增殖和侵襲。Qin等[19]研究顯示,miR-20a過表達會抑制RUNX3表達,進而激活TGF-β信號通路,促進非小細胞肺癌的增殖、遷移和侵襲。在本研究中,PIC促進了Hela細胞中RUNX3表達,提示RUNX3可能參與PIC對Hela細胞抗癌作用。隨后,通過雙熒光素酶報告基因檢測發現,RUNX3可能為miR-106b-5p的靶基因;Western blot證實抑制miR-106b-5p表達可有效促進RUNX3表達,抑制RUNX3表達可逆轉敲低miR-106b-5p表達對Hela細胞的遷移和侵襲的抑制作用。以上結果提示miR-106b-5p可能靶向負調控RUNX3的表達。
綜上所述,PIC可抑制Hela細胞增殖、遷移和侵襲,其分子機制可能與抑制miR-106b-5p表達、促進RUNX3表達有關。本研究表明PIC可能是CC的一種有效治療選擇,為探索CC發展的分子機制研究提供了參考。但本研究僅選擇了一種CC細胞系進行研究,且并未探究PIC調控miR-106b-5p/RUNX3軸抑制CC遷移和侵襲是否還有其他因子參與,在后續的研究中將結合體內動物實驗進行更深入更全面的探索。
參考文獻
[1] SUNG H,FERLAY J,SIEGEL R L,et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA Cancer J Clin,2021,71(3):209-249. doi:10.3322/caac.21660.
[2] LEI J,ARROYO-M?HR L S,LAGHEDEN C,et al. Human papillomavirus infection determines prognosis in cervical cancer[J]. J Clin Oncol,2022,40(14):1522-1528. doi:10.1200/JCO.21.01930.
[3] MAYADEV J S,KE G,MAHANTSHETTY U,et al. Global challenges of radiotherapy for the treatment of locally advanced cervical cancer[J]. Int J Gynecol Cancer,2022,32(3):436-445. doi:10.1136/ijgc-2021-003001.
[4] WANG K J,ZHANG W Q,LIU J J,et al. Piceatannol protects against cerebral ischemia/reperfusion?induced apoptosis and oxidative stress via the Sirt1/FoxO1 signaling pathway[J]. Mol Med Rep,2020,22(6):5399-5411. doi:10.3892/mmr.2020.11618.
[5] WANG B,LI J. Piceatannol suppresses the proliferation and induced apoptosis of osteosarcoma cells through PI3K/AKT/mTOR pathway[J]. Cancer Manag Res,2020,12(3):2631-2640. doi:10.2147/CMAR.S238173.
[6] LI X G,ZHANG W S,YAN X P. Piceatannol inhibits proliferation and induces apoptosis of bladder cancer cells through regulation of the PTEN/AKT signal pathway[J]. Cell Mol Biol (Noisy-le-grand),2020,66(3):181-184.
[7] ALJABALI A A A,BAKSHI H A,HAKKIM F L,et al. Albumin nano-encapsulation of piceatannol enhances its anticancer potential in colon cancer via downregulation of nuclear p65 and HIF-1α[J]. Cancers (Basel),2020,12(1):113-118. doi:10.3390/cancers12010113.
[8] FAN Y,SHENG W,MENG Y,et al. LncRNA PTENP1 inhibits cervical cancer progression by suppressing miR-106b[J]. Artif Cells Nanomed Biotechnol,2020,48(1):393-407. doi:10.1080/21691401.2019.1709852.
[9] GU H,GU S,ZHANG X,et al. miR-106b-5p promotes aggressive progression of hepatocellular carcinoma via targeting RUNX3[J]. Cancer Med,2019,8(15):6756-6767. doi:10.1002/cam4.2511.
[10] 戴云峰. RUNX3和VEGF在宮頸癌組織中的表達分析[J]. 中國醫藥指南,2020,18(9):51-52. DAI Y F. Expression analysis of RUNX3 and VEGF in cervical carcinoma[J]. Guide of China Medicine,2020,18(9):51-52. doi:10.15912/j.cnki.gocm.2020.09.028.
[11] SUN X,FU P,XIE L,et al. Resveratrol inhibits the progression of cervical cancer by suppressing the transcription and expression of HPV E6 and E7 genes[J]. Int J Mol Med,2021,47(1):335-345. doi:10.3892/ijmm.2020.4789.
[12] 王旭,徐文舉,袁五營. miRNA-4262通過靶向神經調節蛋白1調控非小細胞肺癌細胞增殖,侵襲,遷移的作用機制[J]. 癌癥進展,2020,18(2):133-137. WANG X,XU W J,YUAN W Y. Mechanism of miRNA-4262 regulating the proliferation, invasion and migration of non-small cell lung cancer cells by targeting NRG1[J]. Oncology Progress,2020,18(2):133-137. doi:10.11877/j.issn.1672-1535.2020.18.02.07.
[13] KUMAR A, RIMANDO A M, LEVENSON A S. Resveratrol and pterostilbene as a microRNA-mediated chemopreventive and therapeutic strategy in prostate cancer[J]. Ann N Y Acad Sci,2017,1403(1):15-26. doi:10.1111/nyas.13372.
[14] DU M,ZHANG Z,GAO T. Piceatannol induced apoptosis through up-regulation of microRNA-181a in melanoma cells[J]. Biol Res,2017,50(1):36-45. doi:10.1186/s40659-017-0141-8.
[15] PAN M,CHEN Q,LU Y,et al. MiR-106b-5p regulates the migration and invasion of colorectal cancer cells by targeting FAT4[J]. Biosci Rep,2020,40(11):98-105. doi:10.1042/BSR20200098.
[16] YU L X,ZHANG B L,YANG M Y,et al. MicroRNA-106b-5p promotes hepatocellular carcinoma development via modulating FOG2[J]. Onco Targets Ther,2019,12(5):5639-5647. doi:10.2147/OTT.S203382.
[17] XIAO Z,TIAN Y,JIA Y,et al. RUNX3 inhibits the invasion and migration of esophageal squamous cell carcinoma by reversing the epithelial-mesenchymal transition through TGF-β/Smad signaling[J]. Oncol Rep,2020,43(4):1289-1299. doi:10.3892/or.2020.7508.
[18] SONG J,LIU Y,WANG T,et al. MiR-17-5p promotes cellular proliferation and invasiveness by targeting RUNX3 in gastric cancer[J]. Biomed Pharmacother,2020,128(4):110246-110253. doi:10.1016/j.biopha.2020.110246.
[19] QIN X,WANG X Y,FEI J W,et al. MiR-20a promotes lung tumorigenesis by targeting RUNX3 via TGF-β signaling pathway[J]. J Biol Regul Homeost Agents,2020,34(2):12-20. doi:10.23812/20-12A.
(2022-09-22收稿 2023-03-03修回)
(本文編輯 李鵬)

