預擴增qPCR技術檢測少量豬早期胚胎細胞基因表達的研究


摘" 要: 旨在比較3種qPCR方法檢測少量豬早期胚胎細胞多能性和組蛋白乙酰化修飾相關基因的動態表達情況。本研究收集不同時期(1-細胞、2-細胞、4-細胞、8-細胞、桑葚胚和囊胚)豬孤雌激活胚胎,利用常規RT-qPCR、cDNA預擴增qPCR和樣本直接預擴增qPCR檢測微量胚胎細胞多能性和組蛋白乙酰化修飾相關基因的表達情況。結果顯示,預擴增qPCR具有穩定的擴增曲線,熔解曲線呈現穩定單峰,而常規RT-qPCR檢測的擴增曲線循環閾值在35以上,熔解曲線呈現多峰;胚胎細胞直接預擴增后的樣品稀釋20 000倍仍能檢測目的基因的表達,且能穩定檢測單個胚胎細胞的基因表達情況;多能性和組蛋白乙酰化修飾相關基因在豬孤雌激活早期胚胎發育過程中呈現先升高后降低的表達趨勢,在基因組激活階段的表達水平最高。綜上表明,樣本直接預擴增qPCR的檢測靈敏性和準確性更高,且操作相對簡單、成本較低,適用于微量胚胎細胞的基因表達檢測,可為研究早期胚胎發育機制提供方法參考。
關鍵詞: 預擴增;qPCR;豬早期胚胎;微量細胞;基因表達
中圖分類號:S828.3
文獻標志碼:A
文章編號:0366-6964(2024)12-5567-08
doi: 10.11843/j.issn.0366-6964.2024.12.021
開放科學(資源服務)標識碼(OSID):
收稿日期:2024-04-15
基金項目:廣東省基礎與應用基礎研究基金委員會區域聯合基金—青年基金項目(2021A1515110046);深圳市可持續發展科技專項項目(KCXFZ20201221173213037);國家自然科學基金青年項目(32302747)
作者簡介:晏" 超(1999-),男,江西宜春人,碩士生,主要從事動物遺傳育種與繁殖研究,E-mail: 15170529593@163.com
*通信作者:唐中林,主要從事豬重要經濟性狀的遺傳改良研究,E-mail: tangzhonglin@caas.cn;陳指龍,主要從事豬繁殖與胚胎發育研究,E-mail: chenzhilong@caas.cn
Detection of Gene Expression in Trace Cells of Early Porcine Embryo by Pre-amplified
Quantitative PCR
YAN" Chao1,2, LIU" Yonggang2, XIE" Hao1, PENG" Cuiting1, ZHANG" Caiyong1, ZHAO" Yulan1,3, QI" Lin1, CHEN" Zhilong1,3,4,5*, TANG" Zhonglin1,3,4,5*
(1.Agricultural Genomics Institute at Shenzhen, Chinese Academy of Agricultural Sciences, Shenzhen 518000," China;
2.College of Animal Science and Technology, Yunnan Agricultural University, Kunming 650201," China;
3.Kunpeng Institute of Modern Agriculture at Foshan, Foshan 528225," China;
4.Shenzhen Branch Center of Guangdong Laboratory of Lingnan Modern Agricultural Science and Technology, Shenzhen 518000," China;
5.Key Laboratory of Livestock and Poultry Biohistology of Ministry of Agriculture and Rural Affairs, Shenzhen 518000," China)
Abstract: This study aimed to compare 3 quantitative polymerase chain reaction (qPCR) methods for detecting the dynamic expression of genes associated with pluripotency and histone acetylase modification in trace cells of porcine early embryo. Porcine parthenogenetic activated embryos at different stages (1-cell, 2-cell, 4-cell, 8-cell, morula and blastocyst) were collected, and the expression of genes associated with cell pluripotency and histone acetylase modification were detected by conventional RT-qPCR, cDNA pre-amplified qPCR and sample direct pre-amplified qPCR. The results indicated that the pre-amplified qPCR exhibited a stable amplification curve, and the melt curve displayed a consistent single peak, the conventional RT-qPCR produced cyclic thresholds above 35 and multiple peaks in the melt curve. Notably, target gene expression was successfully detected even after a 20 000-fold dilution of embryonic cells using pre-amplification, and gene expression at the single embryonic cells could be reliably assessed. The expression patterns of genes related to pluripotency and histone acetylase modification exhibited an initial increase followed by a decline across different stages of porcine parthenogenetic activated embryos, with the highest expression levels occurring at the genome activation stage. In conclusion, pre-amplified qPCR demonstrates superior sensitivity and accuracy, with a relatively simple operational protocol and lower costs, making it a suitable approach for gene expression analysis in trace cells of embryo. This methodology has the potential to advance the understanding for the mechanisms underlying early embryonic development.
Key words: preamplification; qPCR; porcine early embryo; trace cells; gene expression
*Corresponding authors:" TANG Zhonglin, E-mail: tangzhonglin@caas.cn; CHEN Zhilong, E-mail: chenzhilong@caas.cn
豬早期胚胎發育過程涉及復雜的基因轉錄變化和表觀重塑,包括母源RNA降解和合子基因組激活,伴隨著細胞由全能性向多能性的轉變,關鍵基因的特定時空表達是確保胚胎正常發育的基礎[1-3]。基因轉錄豐度的檢測有助于判斷早期胚胎的發育潛能和解析發育機制,為后續提高體外生產胚胎的發育率奠定基礎[4,5]。
實時定量PCR(real-time quantitative PCR, qPCR)和RNA測序(RNA sequencing, RNA-seq)技術是檢測基因表達譜的常規方法。由于胚胎細胞數量有限,基因轉錄豐度較低,利用低輸入RNA-seq需經過多輪全轉錄組擴增后進行測序分析,費時又費力,且容易出現全轉錄組擴增偏差,比實時定量PCR的靈敏度更低,變化更大[6,7]。qPCR檢測表達豐度低的基因需要足夠數量的模板,且多基因檢測對模板數量的要求更高[8]。預擴增檢測技術可通過增加樣本特異性核苷酸序列的拷貝數,實現低表達豐度目的基因被高效檢測,廣泛應用于人類植入前胚胎遺傳學檢測、哺乳動物性別鑒定、胚胎基因組選擇育種等領域[9-13]。核酸特異性預擴增反應使用低濃度引物(比常規PCR低10~20倍),在有限的循環次數(20個循環或更短)內延長退火時間(3 min或更長時間),確保所有待測核酸穩定形成特異性PCR擴增產物;此外,預擴增反應適用于大批量核酸檢測(引物庫≥96 次),檢測效率比常規qPCR高[14]。本研究比較了常規反轉錄qPCR(reverse transcription-qPCR, RT-qPCR)和預擴增qPCR檢測少量豬早期胚胎細胞基因表達的準確性和靈敏性,揭示了多能性和組蛋白乙酰化修飾相關基因在豬孤雌激活胚胎發育過程中的動態表達水平,為進一步研究豬早期胚胎發育機制奠定基礎。
1" 材料與方法
1.1" 試驗材料與試劑
豬卵巢來源于當地屠宰場。孕馬血清促性腺激素和人絨毛膜促性腺激素購于ProSpec生物公司,TCM-199粉末、胎牛血清(fetal bovine serum, FBS)、杜氏磷酸鹽緩沖液(dulbecco’s phosphate buffered saline, DPBS)和青-鏈霉素購于Gibco公司,微量RNA提取試劑盒購于天根生化科技有限公司,SYBR qPCR Master Mix、單細胞特異序列擴增試劑盒和cDNA反轉錄試劑盒購于諾唯贊生物公司,其他試劑無特殊說明外均購于Sigma公司。
1.2" 豬卵母細胞的分離與成熟培養
屠宰場采集的卵母細胞用33℃恒溫箱在4 h內運送回實驗室,用含2%青-鏈霉素的生理鹽水清洗后,用刀片劃破2~6 mm的卵泡,收集卵泡液至15 mL圓底管。卵泡液靜置15 min后,去除上清,用清洗液(含3% FBS的DPBS)清洗一遍,再加入體外操作液(含3%牛血清白蛋白的TCM-199溶液),顯微鏡下挑取形態飽滿、含3層致密卵丘細胞包裹的卵丘-卵母細胞復合體(COCs),每50個COCs放入含400 μL卵母細胞體外成熟液[15]的4孔板中,置于38.5 ℃、5% CO2培養箱中培養42~44 h。
1.3" 豬卵母細胞孤雌激活胚胎的收集
體外成熟后的豬卵母細胞用含1 mg·mL-1 透明質酸酶的TCM-199液吹打脫落卵丘細胞,挑取形態飽滿、極體明顯的卵母細胞至T2(含2% FBS的TCM-199液)液滴中清洗3次,移入激活液(含0.05 mmol·L-1 CaCl2,0.1 mmol·L-1 MgSO4,0.3 mol·L-1 甘露醇,1 mg·mL-1 聚乙烯醇)中平衡3 min,置于融合槽電極中間,以200 V·mm-1、30 μs·次-1電激活1次,T20(含20%FBS的TCM-199液)清洗3次再轉移至含5 mg·mL-1 細胞松弛素B和10 mg·mL-1 環己亞胺的PZM-3胚胎體外培養基中[16],置于38.5 ℃、5% CO2、5% O2和90% N2培養箱中進行培養。
收集豬MII期卵母細胞和不同發育時期豬孤雌激活胚胎(1-細胞、2-細胞、4-細胞、8-細胞、桑葚胚和囊胚),先用含有0.1%聚乙烯醇的DPBS溶液清洗3遍,再用酸性Tyrode’s液去除透明帶,清洗液清洗后將每組樣本裝入含350 μL裂解液(來源于RNA提取試劑盒)的1.5 mL無菌無核酸酶的離心管中,用于微量RNA提取。預擴增胚胎細胞樣本則在去除透明帶后移入200 μL無菌無核酸酶的空離心管中,迅速放入液氮中,用于預擴增qPCR。
1.4" 實時熒光定量PCR
將收集的胚胎樣本分以下3組進行試驗:1)常規RT-qPCR。根據微量RNA提取試劑盒說明書操作步驟提取胚胎細胞樣本(25個細胞)總RNA,用反轉錄試劑盒將mRNA反轉錄成cDNA。2)cDNA預擴增qPCR。將上述反轉錄的cDNA樣本進行預擴增反應,10 μL體系包含5 μL 2×反應MIX、0.2 μL RT/Taq酶、2.8 μL無核酸酶水、0.1 μmol·L-1引物混合液和1 μL cDNA樣本。先將反應混合液置于液氮2 min,3 000 r·min-1 離心2 min,再在PCR儀上95 ℃ 3 min,17個循環(95 ℃ 15 s,60 ℃ 4 min),加入5 μL Exonuclease I(1 U·μL-1),PCR儀中37℃反應30 min,85℃反應15 min去除引物。3)直接預擴增qPCR。將豬胚胎細胞用0.5 μL DNase I(1 U·μL-1)在37℃反應15 min去除基因組DNA后進行預擴增反應。15 μL體系包含7.5 μL 2×反應MIX、0.3 μL RT/Taq酶、4 μL無核酸酶水、0.1 μmol·L-1 引物混合液和1.7 μL樣本。先將反應混合液置于液氮2 min,3 000 r·min-1 離心2 min,再到PCR儀上50℃ 60 min,95℃ 3 min,17個循環(95 ℃ 15 s,60 ℃ 4 min),用Exonuclease I去除引物。
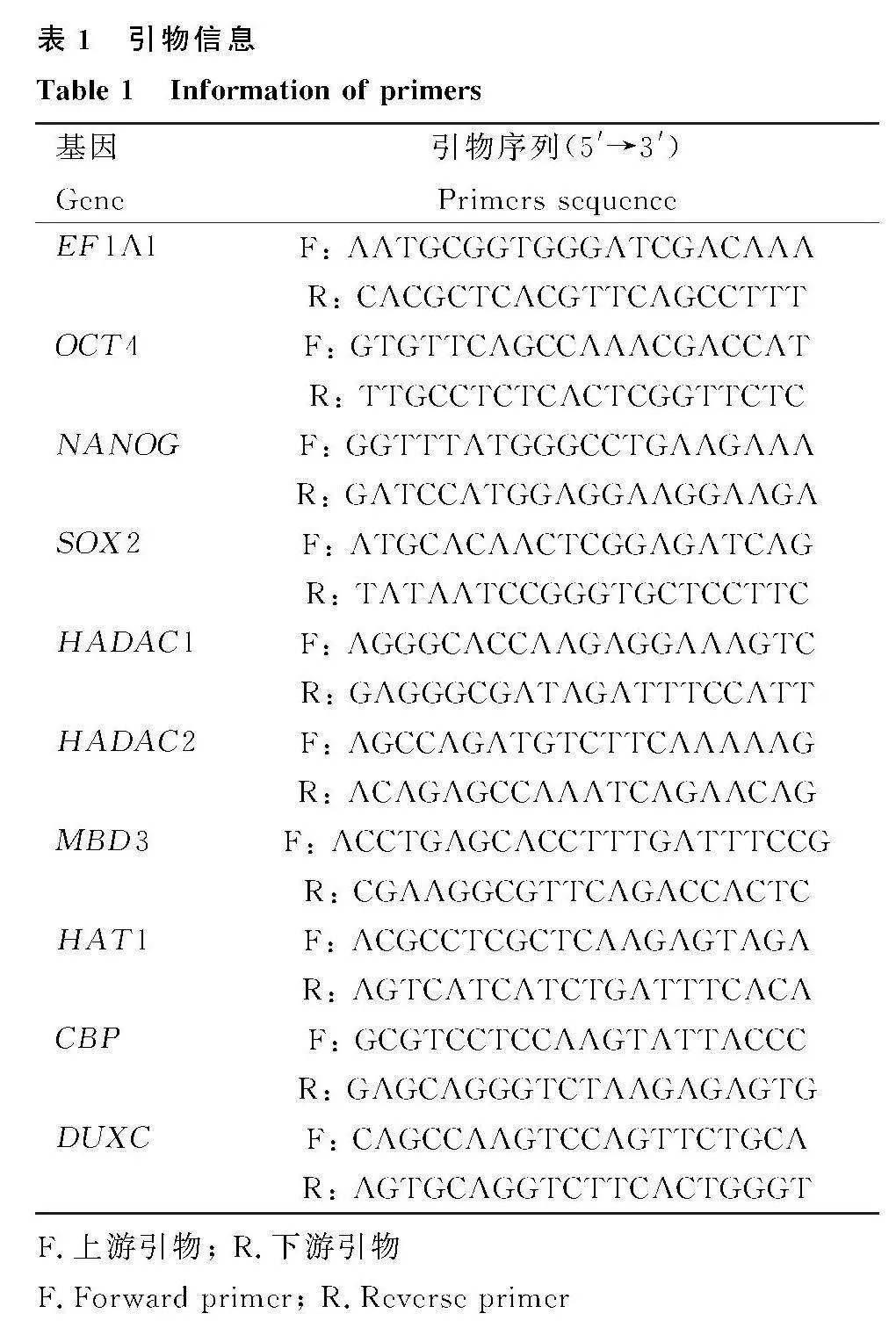
將上述3種樣本用SYBR qPCR Master Mix試劑盒進行qPCR檢測。10 μL反應體系:2×SYBR Mix 5 μL,上、下游引物各0.2 μL,樣本1 μL,無核酸酶水3.6 μL。引物信息見表1。反應條件:95 ℃預變性30 s,40個循環(95 ℃變性10 s,60 ℃退火延伸30 s)。熔解曲線:95 ℃ 15 s , 60 ℃ 60 s, 95 ℃ 15 s。目的基因表達水平對比內參基因后采用2-ΔΔCt法計算結果。
1.5" 數據的處理與分析
上述試驗獨立重復4次,結果利用GraphPad Prism軟件繪制數據圖例。
2" 結" 果
2.1" 三種qPCR方法的檢測靈敏性
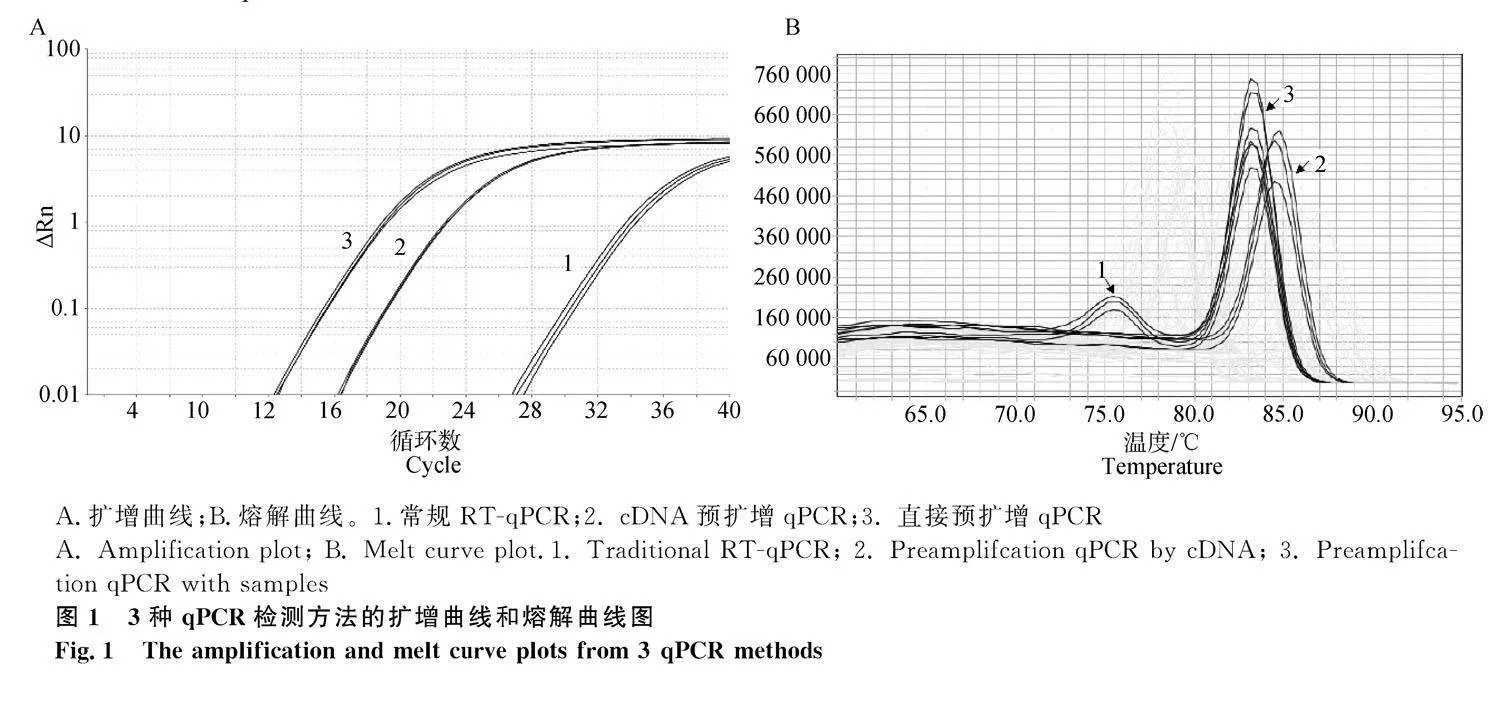
采用3種qPCR方法檢測25個豬MⅡ期卵母細胞的內參基因EF1A1表達情況。結果如圖1所示,常規RT-qPCR的擴增曲線CT在35以上,且熔解曲線出現多峰,cDNA和直接預擴增qPCR的擴增曲線CT在30以內,熔解曲線呈現穩定的單峰。說明預擴增qPCR的靈敏性高于常規RT-qPCR。
2.2" 不同樣本量的預擴增qPCR
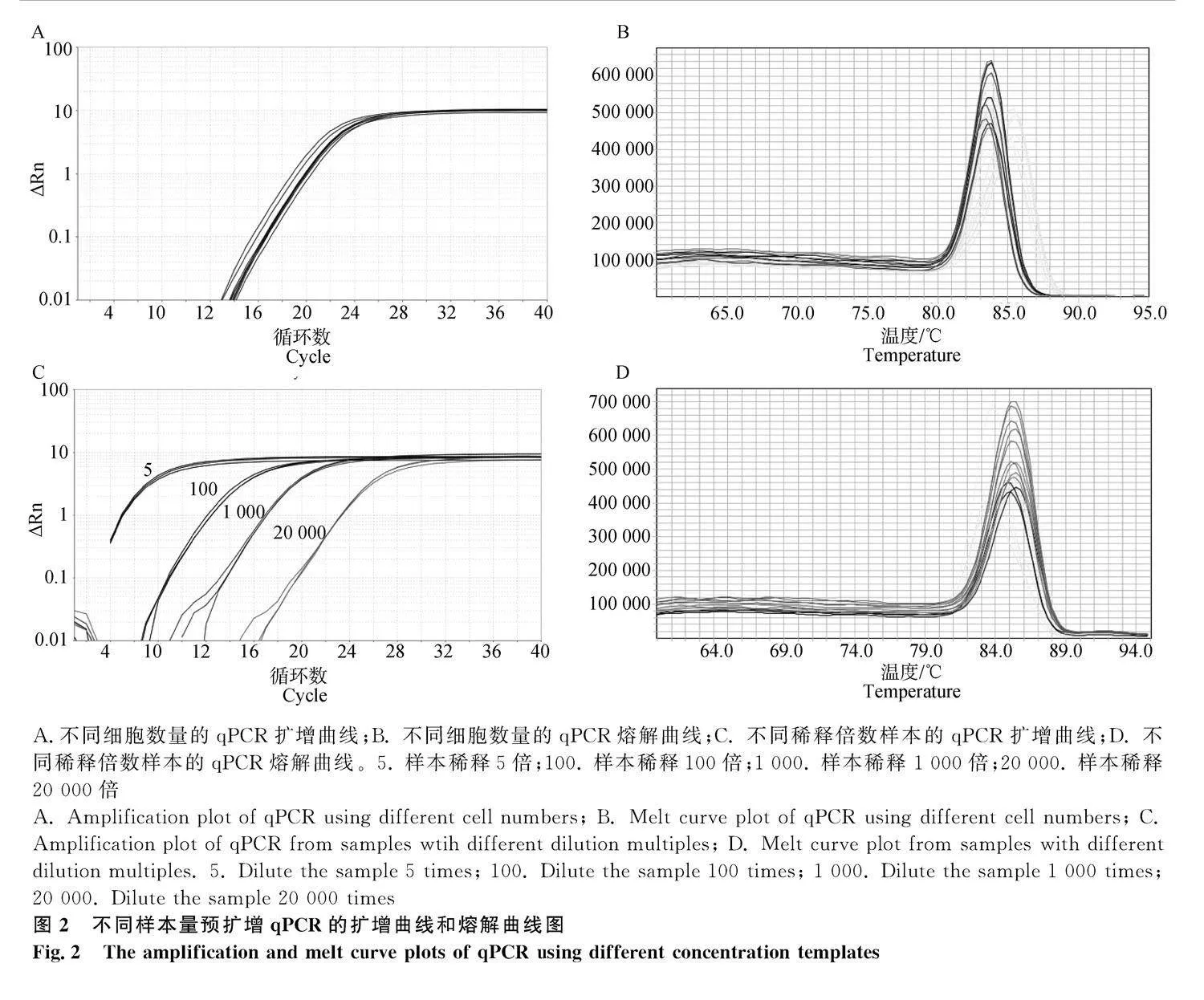
用直接預擴增qPCR分別檢測1個、5個和25個MⅡ期卵母細胞的EF1A1基因表達情況。結果如圖2A和2B所示,3組細胞樣本的擴增曲線CT值均在22~24之間,熔解曲線呈現穩定單峰。進一步對25個MⅡ期卵母細胞的預擴增后樣本進行5、100、1 000和20 000倍稀釋。結果顯示,隨著稀釋倍數的增加,CT值相應增加,保持在28之內,熔解曲線呈現穩定的單峰(圖2C和2D)。結果說明,直接預擴增qPCR可檢測單個細胞樣本的基因表達,細胞樣本稀釋20 000倍后仍能有效檢測目的基因的表達。
2.3" 三種qPCR檢測方法的準確性比較
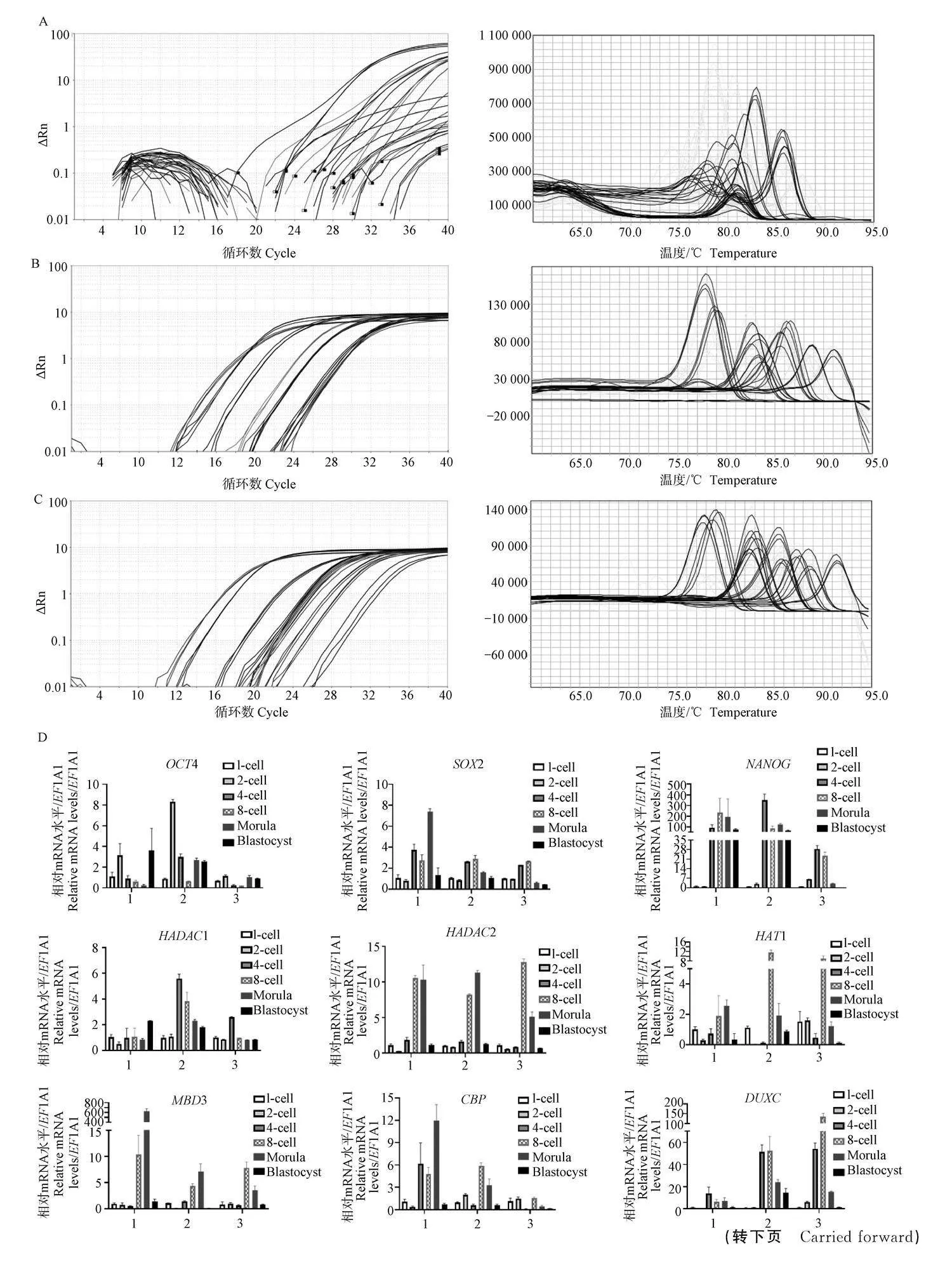
利用3種qPCR方法檢測不同發育時期豬孤雌激活胚胎細胞(25個1-細胞、20個2-細胞、15個4-細胞、12個8-細胞、10個桑葚胚和10個囊胚)中多能性和組蛋白修飾相關基因的表達情況。結果如圖3所示,常規RT-qPCR檢測目的基因的CT值均在35以上,且熔解曲線出現多峰,而兩種預擴增qPCR的CT明顯下降,熔解曲線呈現穩定的單峰。兩種預擴增qPCR檢測的多能性和組蛋白修飾相關基因表達趨勢較為一致,基因表達檢測結果的組內誤差較低。綜上說明,預擴增qPCR的檢測準確性要優于常規RT-qPCR。
3" 討" 論
隨著下一代測序(二代)技術的發展,微量細胞測序技術陸續被開發并廣泛應用于胚胎發育機制研究[17-19]。RNA-seq和RT-qPCR是檢測微量細胞樣本mRNA水平的常用方法[11,20]。對于特定基因或基因集的表達檢測,qPCR相比RNA-seq操作更簡便、成本較低且具有更高的靈敏性和準確性[9,21]。預擴增qPCR可在進行PCR反應前通過增加樣本中特異性核苷酸序列的拷貝數來檢測有限核酸數量樣本的基因表達水平[22]。早在2010年,Hernndez-Arteage和Lpez-Revilia[23]應用預擴增qPCR技術,將樣本稀釋50倍后進行15個循環數的預擴增反應,可使樣本中癌基因表達檢測的靈敏度提高10 750倍。Ankica等[10]利用預擴增qPCR技術高效檢測了血液中微量Micro RNAs的表達情況。此外,Xiao等[24]利用預擴增qPCR技術確定了單個牛囊胚的性別和性染色體相關基因的表達情況。本研究通過3種qPCR方法檢測不同發育時期豬胚胎多能性和組蛋白乙酰化修飾相關基因的表達水平,發現預擴增qPCR的檢測靈敏性高于傳統
RT-qPCR,傳統RT-qPCR無法有效檢測少量細胞中目的基因的表達,擴增曲線和熔解曲線均不穩定,且組內誤差較大。預擴增反應能有效增加樣本模板數量,同時可減少樣品提取核酸過程造成的組別差異,但有限數量的胚胎細胞樣本很難估算RNA濃度,只能通過擴增曲線和熔解曲線反應預擴增效果[25,26]。Kroneis等[9]研究表明,24個擴增循環數足夠使30 pg RNA預擴增成cDNA。本研究在前期摸索了不同循環數對25個豬卵母細胞的預擴增效果,發現17個循環數進行預擴增反應能有效檢測20個基因的表達水平。由于收集的胚胎細胞數量有限,在某些特定情況下,可能需要對極少量甚至1個胚胎或者細胞進行測定。Xiao等[11]將單個囊胚預擴增后的cDNA樣本稀釋1 024倍,還可有效檢測目的基因的表達。本研究對預擴增樣本稀釋20 000倍后仍能有效檢測到EF1A1基因的表達,根據擴增曲線CT值,25個細胞預擴增后稀釋1 000倍進行qPCR較為合適。此外,1個、5個和25個細胞進行預擴增qPCR,其CT值相近,說明預擴增反應可穩定檢測單個細胞中基因表達,且25個之內細胞數進行預擴增后的cDNA含量相近。
哺乳動物早期胚胎發育經歷了合子基因組激活和細胞由全能性向多能性轉變的過程,組蛋白乙酰化修飾在胚胎合子基因組激活過程中起著重要的調控作用[27,28],先鋒轉錄因子OCT4、SOX2和NANOG是調控胚胎細胞多能性的重要基因[29-31]。本研究通過預擴增qPCR檢測發現,豬孤雌激活胚胎多能性和組蛋白乙酰化修飾基因SOX2、NANOG、HADAC1、HADAC2、HAT1、MBD3、CBP和DUXC在合子基因組激活階段(4-~8-細胞時期)呈現高表達,與Li等[32]的報道基本一致。豬胚胎4-~8-細胞階段母源mRNA不斷降解,合子基因被激活,多能性和組蛋白乙酰化修飾基因特定時空的動態表達是確保胚胎正常發育的基礎[33-36]。
cDNA預擴增qPCR檢測豬胚胎多能性和組蛋白乙酰化修飾基因的表達趨勢與預擴增qPCR相似,但cDNA預擴增qPCR的目的基因表達水平相對較高,可能是cDNA預擴增樣本殘留RNA導致。綜上,樣本直接預擴增qPCR的檢測準確性更高。
4" 結" 論
本研究優化的樣本直接預擴增qPCR技術比常規RT-qPCR和cDNA預擴增qPCR具有更高的準確性和靈敏性。通過少量胚胎細胞樣本直接預擴增qPCR技術揭示了不同發育時期豬孤雌激活胚胎多能性和組蛋白乙酰化修飾相關基因的動態表達水平,呈現先升高后降低的表達趨勢,在合子基因組激活階段的表達水平最高。本研究可為進一步解析豬早期胚胎發育機制提供參考。
參考文獻(References):
[1]" ZHU W,BU G W,HU R F,et al.KLF4 facilitates chromatin accessibility remodeling in porcine early embryos[J].Sci China Life Sci,2024,67(1):96-112.
[2]" BU G W,ZHU W,LIU X,et al.Coordination of zygotic genome activation entry and exit by H3K4me3 and H3K27me3 in porcine early embryos[J].Genome Res,2022,32(8):1487-1501.
[3]" HE T Y,PENG J Y,YANG S,et al.SINE-associated LncRNA SAWPA regulates porcine zygotic genome activation[J].Adv Sci (Weinh),2024,11(2):2307505.
[4]" GAO R N,LI Q C,QIU M Y,et al.Serum exosomal miR-192 serves as a potential detective biomarker for early pregnancy screening in sows[J].Anim Biosci,2023,36(9):1336-1349.
[5]" ZOLINI A M,BLOCK J,RABAGLINO M B,et al.Genes associated with survival of female bovine blastocysts produced in vivo[J].Cell Tissue Res,2020,382(3):665-678.
[6]" GROFF A F,RESETKOVA N,DIDOMENICO F,et al.RNA-seq as a tool for evaluating human embryo competence[J].Genome Res,2019,29(10):1705-1718.
[7]" LUO Q K,ZHANG H.Emergence of bias during the synthesis and amplification of cDNA for scRNA-seq[J].Adv Exp Med Biol,2018,1068:149-158.
[8]" STHLBERG A,KUBISTA M.The workflow of single-cell expression profiling using quantitative real-time PCR[J].Expert Rev Mol Diagn,2014,14(3):323-331.
[9]" KRONEIS T,JONASSON E,ANDERSSON D,et al.Global preamplification simplifies targeted mRNA quantification[J].Sci Rep,2017,7:45219.
[10]" SEKOVANIC′ A,DOROTIC′ A,JURASOVIC′ J,et al.Pre-amplification as a method for improvement of quantitative RT-PCR analysis of circulating miRNAs[J].Biochem Med (Zagreb),2021,31(1):010901.
[11]" XIAO Y,SOSA F,DE ARMAS L R,et al.An improved method for specific-target preamplification PCR analysis of single blastocysts useful for embryo sexing and high-throughput gene expression analysis[J].J Dairy Sci,2021,104(3):3722-3735.
[12]" MOGHADDASZADEH-AHRABI S,FARAJNIA S,RAHIMI-MIANJI G,et al.A short and simple improved-primer extension preamplification (I-PEP) procedure for whole genome amplification (WGA) of bovine cells[J].Anim Biotechnol,2012,23(1):24-42.
[13]" SUN S Q,ABOELENAIN M,ARIAD D,et al.Identifying risk variants for embryo aneuploidy using ultra-low coverage whole-genome sequencing from preimplantation genetic testing[J].Am J Hum Genet,2023,110(12):2092-2102.
[14]" ANDERSSON D,AKRAP N,SVEC D,et al.Properties of targeted preamplification in DNA and cDNA quantification[J].Expert Rev Mol Diagn,2015,15(8):1085-1100.
[15]" YUAN Y,SPATE L D,REDEL B K,et al.Quadrupling efficiency in production of genetically modified pigs through improved oocyte maturation[J].Proc Natl Acad Sci U S A,2017,114(29):E5796-E5804.
[16]" YOSHIOKA K,SUZUKI C,TANAKA A,et al.Birth of piglets derived from porcine zygotes cultured in a chemically defined medium[J].Biol Reprod,2002,66(1):112-119.
[17]" ZHOU F,WANG R,YUAN P,et al.Reconstituting the transcriptome and DNA methylome landscapes of human implantation[J].Nature,2019,572(7771):660-664.
[18]" WU Y,XU X C,QI M J,et al.N6-methyladenosine regulates maternal RNA maintenance in oocytes and timely RNA decay during mouse maternal-to-zygotic transition[J].Nat Cell Biol,2022,24(6):917-927.
[19]" XU R M,ZHU Q S,ZHAO Y Y,et al.Unreprogrammed H3K9me3 prevents minor zygotic genome activation and lineage commitment in SCNT embryos[J].Nat Commun,2023,14(1):4807.
[20]" LAVAGI I,KREBS S,SIMMET K,et al.Single-cell RNA sequencing reveals developmental heterogeneity of blastomeres during major genome activation in bovine embryos[J].Sci Rep,2018,8(1):4071.
[21]" KOLODZIEJCZYK A A,LNNBERG T.Global and targeted approaches to single-cell transcriptome characterization[J].Brief Funct Genomics,2018,17(4):209-219.
[22]" KORENKOV V,SCOTT J,NOVOSADOV V,et al.Pre-amplification in the context of high-throughput qPCR gene expression experiment[J].BMC Mol Biol,2015,16:5.
[23]" HERNNDEZ-ARTEAGA S,LPEZ-REVILLA R.Ultrasensitive quantitation of human papillomavirus type 16 E6 oncogene sequences by nested real time PCR[J].Infect Agent Cancer,2010,5:9.
[24]" XIAO Y,SOSA F,DE ARMAS L R,et al.An improved method for specific-target preamplification PCR analysis of single blastocysts useful for embryo sexing and high-throughput gene expression analysis[J].J Dairy Sci,2021,104(3):3722-3735.
[25]" ANDERSSON D,AKRAP N,SVEC D,et al.Properties of targeted preamplification in DNA and cDNA quantification[J].Expert Rev Mol Diagn,2015,15(8):1085-1100.
[26]" JANNAMAN E A,XIAO Y,HANSEN P J.Actions of colony-stimulating factor 3 on the maturing oocyte and developing embryo in cattle[J].J Anim Sci,2020,98(4):skaa115.
[27]" WU K L,FAN D D,ZHAO H,et al.Dynamics of histone acetylation during human early embryogenesis[J].Cell Discov,2023, 9(1):29.
[28]" YANG G,ZHANG L F,LIU W Q,et al.Dux-mediated corrections of aberrant H3K9ac during 2-cell genome activation optimize efficiency of somatic cell nuclear transfer[J].Cell Stem Cell,2021,28(1):150-163.e5.
[29]" LI L J,LAI F N,HU X Y,et al.Multifaceted SOX2-chromatin interaction underpins pluripotency progression in early embryos[J].Science,2023,382(6676):eadi5516.
[30]" STIRPARO G G,KUROWSKI A,YANAGIDA A,et al.OCT4 induces embryonic pluripotency via STAT3 signaling and metabolic mechanisms[J].Proc Natl Acad Sci U S A,2021,118(3):e2008890118.
[31]" LAI F N,LI L J,HU X Y,et al.NR5A2 connects zygotic genome activation to the first lineage segregation in totipotent embryos[J].Cell Res,2023,33(12):952-966.
[32]" LI X,ZOU C,LI M X,et al.Transcriptome analysis of in vitro fertilization and parthenogenesis activation during early embryonic development in pigs[J].Genes (Basel),2021,12(10):1461.
[33]" WEI Q Q,LI R Q,ZHONG L,et al.Lineage specification revealed by single-cell gene expression analysis in porcine preimplantation embryos[J].Biol Reprod,2018,99(2):283-292.
[34]" KONG Q R,YANG X,ZHANG H,et al.Lineage specification and pluripotency revealed by transcriptome analysis from oocyte to blastocyst in pig[J].FASEB J,2020,34(1):691-705.
[35]" LEE M,CHOI K H,OH J N,et al.SOX2 plays a crucial role in cell proliferation and lineage segregation during porcine pre-implantation embryo development[J].Cell Prolif,2021,54(8):e13097.
[36]" DE MACEDO M P,GLANZNER W G,GUTIERREZ K,et al.Simultaneous inhibition of histone deacetylases and RNA synthesis enables totipotency reprogramming in pig SCNT embryos[J].Int J Mol Sci,2022,23(22):14142.
(編輯" 郭云雁)

